econometron.Models.dynamicsge.linear_dsge
classin theeconometron.Models.dynamicsgemodule
Overview
A Dynamic Stochastic General Equilibrium (DSGE) model describes the decisions of households, firms, and policymakers under uncertainty, subject to market-clearing conditions. A linear DSGE model is the first-order Taylor approximation of the model’s nonlinear equilibrium conditions around a deterministic steady state. The linear_dsge class in econometron.Models.dynamicsge allows specification, solution, and analysis of linear and log-linear DSGE models.
Linearization
Given a nonlinear equilibrium system:
let be the steady state. The linearized system is obtained by a first-order expansion in deviations:
Example (levels) If:
then, around :
- Units: original scale
- Use case: variables can be negative (e.g., interest rate gaps)
Log-Linearization
For positive variables, define the log-deviation:
Apply the Taylor expansion to the logged equations.
Example (logs) If:
then:
- Units: percentage deviations from steady state
- Use case: real, strictly positive variables (e.g., output, capital)
In Practice
- Linearization: use for rates, spreads, gaps.
- Log-linearization: use for real quantities.
class linear_dsge capabilities
- Parse string-based equations
- Compute steady states
- Linearize or log-linearize
- Solve RE systems (Klein(2000))
- Simulate and compute IRFs
- Support for analytical and numerical steady-state and derivative computation
Class Definition
from econometron.Models.dynamicsge import linear_dsge
lineardsge=linear_dsge(
equations=list[str],
variables=list[str],
exo_states=list[str],
endo_states=list[str],
parameters=dict[str, float],
approximation=str,
normalize=bool,
shocks=list[str])Parameters
| Name | Type | Description | Default |
|---|---|---|---|
equations | list[str] | Model equations as strings, must include = and time subscripts (_t, _tp1, _tm1). | None |
variables | list[str] | Variable names (e.g., ['A', 'K', 'C']). | None |
exo_states | list[str] | Exogenous state variables. | None |
endo_states | list[str] | Endogenous state variables. | None |
parameters | dict | Model parameters, e.g., {'sigma': 1.5}. | None |
approximation | str | 'linear' or 'log_linear'. | 'linear' |
normalize | dict | Steady-state normalization values. | {} |
shocks | list[str] | Shock variable names. | None |
Attributes
| Attribute | Type | Description |
|---|---|---|
equations_list | list[str] | Provided equations. |
variables | list[str] | All model variables. |
exo_states | list[str] | Exogenous state variables. |
endo_states | list[str] | Endogenous state variables. |
states | list[str] | Combined exogenous + endogenous states. |
controls | list[str] | Variables not in states. |
parameters | dict | Model parameters dictionary. |
approximation | str | Approximation type. |
normalize | dict | Normalization values. |
shocks | list[str] | Shock variables. |
n_vars, n_equations, n_states, n_exo_states, n_endo_states, n_controls | int | Model dimension counts. |
steady_state | pd.Series | Steady-state values. |
A, B, C | np.ndarray | Jacobian matrices. |
f, p | np.ndarray | Policy and state transition matrices. |
irfs | dict[pd.DataFrame] | Impulse response functions. |
simulated | pd.DataFrame | Simulated data. |
initial_guess | numpy array | Initial guess for steady state. |
Methods
1 - validate_entries()
Purpose: Validates inputs during initialization. It checks:
- Non-empty list of equations containing
=. - Non-empty variables list without duplicates.
- Non-empty parameters dictionary.
- Correct data types for states and shocks.
- Non-empty list of equations containing
Returns:
Trueif valid; otherwise raisesValueErroror issues warnings.
2 - set_new_parameters(params)
Purpose:
Updates model parameters (
self.parameters).Returns:
Prints updated parameters or warns if update fails.
3 - set_initial_guess(initial_guess)
Purpose:
Sets initial guess for steady-state computation.
- If approximation="linear", the exogenous shock is set to 0.
- If approximation="log-linear", the exogenous shock is set to 1.
- You can use normalize = {var: value} to assign a specific value to a given variable.Generally, you don’t need to provide this unless required.
- If
set_initial_guessandguessis not specified, it defaults to an array of ones with length equal to the number of endogenous variables.
Returns:
- Stores as
np.arraycould be checked by :- Example
python lineardsge.initial_guess
- Example
- Raises
ValueErrorif size/type mismatch.
- Stores as
4 - compute_ss(guess=None, method='fsolve', options=None)
purpose
- Computes steady state via numerical optimization,currently suppoting
fsolve. if user didn't specify the initial viaguessorset_initial_guess(),it will default to an array of ones with the same length as the endogenous variables.
Parameters:
guess– Initial guess (defaults to ones).method– Only'fsolve'supported.options– Optimizer options.
Returns:
pd.Seriessteady-state values
5 - approximate(method=None, debug=False)
Purpose:
- Approximates the Rational Expectations (RE) model around its steady state using either analytical or numerical Jacobians.
- Uses
_Analytical_jacobians(symbolic, more stable and parsimonious) or_Numerical_jacobians. - Both functions internally call
_reorder_variables, which reorders variables in the sequence: exo_states → endo_states → controls. - Stores Jacobians as
self.A,self.B,self.C. - Once approximated,The model attribute
approximated = Trueand the model can be solved using the Jacobians. - Note : This method must be called after
compute_ss(). Ifapproximate()is called whenapproximated = True, aWarningis raised to notify that the model is already approximated; users should only call this method once after computing the steady state.
Parameters
method:analyticalornumericaldefault :analyticaldebug:bool, optional- If
True, enables debug mode for more verbose output.
- If
Returns
tuple: (A, B, C)- A – Jacobian matrix associated with expectations
- B – Jacobian matrix associated with current endogenous variables
- C – Jacobian matrix associated with exogenous shocks
6 - solve_RE_model(Parameters=None, debug=False)
Purpose:
Solves the Rational Expectations (RE) model of the form
using Klein’s method (generalized Schur decomposition - Solab.m 2000).
- If
Parametersare provided, the Jacobians are recomputed analytically. - Otherwise, it uses the already stored approximated Jacobians.
- The solution is stored in
self.f(policy function) andself.p(state transition function). - This function can be used once The model is approximated.
- If
Parameters
Parameters(dict, optional): Model parameters to override.debug(bool, optional): If True, enables debug mode for more verbose output.
Returns
tuple: (F, P)F – Control function, mapping states to controls:
P – State transition function, governing the law of motion:
7 - _compute_irfs(T=51, t0=1, shocks=None, center=True, normalize=True)
Purpose:
- Computes impulse response functions (IRFs) for a DSGE model, adjusted to match the behavior of the impulse method. IRFs describe the dynamic response of model variables to exogenous shocks.
- Simulates the model’s response to specified shocks over T periods.
- Stores results in self.irfs as a dictionary of DataFrames, with each key corresponding to a shock and its associated IRFs.
- Ensures compatibility with log-linear or linear approximations, with options to
center(return deviations) ornormalize(divide by steady state).
Parameters:
T (int, optional): Number of periods for IRF computation (default: 51).t0 (int, optional): Time period when shocks are applied (default: 1).shocks (dict, optional): Dictionary of shock names and magnitudes. If None, applies a 0.01 shock to each defined shock (default: None).center (bool, optional): If True, returns deviations from steady state; if False, returns levels or log levels (default: True).normalize (bool, optional): If True, normalizes linear approximation IRFs by steady state values(default: True).
Returns:
dict: Dictionary where keys are shock names and values arepd.DataFrameobjects containing IRFs for states, controls, and the shock itself.Notes:
- Requires
f(policy function) andp(state transition function) to be defined. - Raises
ValueErrorif model matrices or shocks are undefined. - Warns if normalization is requested but steady state contains zeros, setting
normalizetoFalse.
- Requires
8 - plot_irfs(shock_names=None, T=41, scale=100, figsize=(12, 4), lw=5, alpha=0.5, title_prefix="IRF", ylabel="Percentage Deviation")
Purpose:
Compute impulse Responses via
_compute_irfs()which are stored inself.irfs.Plots impulse response functions (IRFs) for specified shocks, with separate subplots for the corresponding exogenous state and other variables.
Generates a two-panel plot for each shock: one for the exogenous state,
exo_states, and one for all other variables (states and controls).Scales IRF values for better visualization and allows customization of plot aesthetics.
Parameters:
shock_names (list, optional): List of shocks to plot. If None, plots all shocks in self.irfs (default: None).T (int, optional): Number of periods for IRF computation if not already computed(default: 41).scale (float, optional): Scaling factor for IRF values(default: 100).figsize (tuple, optional): Figure size as (width, height) in inches(default: (12, 4)).lw (float, optional): Line width for plots (default: 5).alpha (float, optional): Line transparency (default: 0.5).title_prefix (str, optional): Prefix for subplot titles (default: "IRF").ylabel (str, optional): Y-axis label (default: "Percentage Deviation").
Returns:
None: Displays the plots for each shock. Notes:- Raises
ValueErrorifself.irfsis empty, not a dictionary, or if specified shocks/columns are missing. - Automatically adjusts y-axis limits for clarity.
- Raises
9 - simulate(T=51, drop_first=300, covariance_matrix=None, seed=None, center=True, normalize=True)
Purpose:
- Simulates the DSGE model dynamics over
Tperiods, adjusted to match the behavior ofstoch_sim. Generates stochastic paths for states and controls driven by exogenous shocks. - Discards
drop_firstinitial periods to eliminate transient dynamics. - Supports custom covariance matrices for shocks or defaults to a diagonal matrix based on shock variances.
- Stores results in
self.simulatedas aDataFrame.
- Simulates the DSGE model dynamics over
Parameters:
T (int, optional): Number of periods to simulate (default: 51).drop_first (int, optional): Number of initial periods to discard (default: 300).covariance_matrix (array-like, optional): Covariance matrix for shocks(n_shocks x n_shocks). If None, uses a diagonal matrix with variances of0.01²(default: None).seed (int, optional): Random seed for reproducibility (default: None).center (bool, optional): IfTrue, returns deviations; if False, returns levels or log levels (default: True).normalize (bool, optional): IfTrue, normalizes linear approximation simulations by steady state (default: True).
Returns:
pd.DataFrame: DataFrame containing simulated shocks and variables (states and controls).Notes:
- Requires
self.fandself.pto be defined (model must be solved). - Raises
ValueErrorif model matrices are undefined or covariance matrix has incorrect dimensions. - Warns if normalization is requested but steady state contains zeros, setting
normalizetoFalse.
- Requires
10 - simulations(title="The Model Simulation", figsize=(12, 8), save_path=None)
- Purpose:
Plots the simulated data from simulate, with exogenous states in separate subplots and endogenous variables (states and controls) together in one subplot.
Creates a multi-panel figure where each exogenous state has its own subplot, and all endogenous variables are plotted together.
Allows saving the plot to a file or displaying it.
Parameters:
title (str, optional): Title of the plot (default: "The Model Simulation").figsize (tuple, optional): Figure size as (width, height) in inches (default: (12, 8)).save_path (str, optional): File path to save the plot (e.g., 'plot.png'). IfNone, displays the plot (default: None).Returns:
None: Displays or saves the plot.- Notes:
- Requires
self.simulatedto be apd.DataFramefrom a prior call tosimulate. - Raises
ValueErrorifself.simulatedis not defined or no exogenous/endogenous variables are found. - Automatically adjusts layout to accommodate the title and subplots.
- Requires
Usage Example – RBC Model
1 We Start By importing The class linear_dsge from the econometron.Models.dynamicsge module.
from econometron.Models.dynamicsge import linear_dsge2 We Define The Model Equations as Follow ,specify all the variables, the exogenous , endogenous states and shocks along with the parameters.
equations = [
"-C_t**(-sigma) + beta * C_tp1**(-sigma) * (alpha * A_tp1 * K_tp1**(alpha-1) + 1 - delta) = 0",
"C_t + K_tp1 - A_t * K_t**alpha - (1 - delta) * K_t = 0",
"-log(A_tp1) + rho_a * log(A_t) + epsilon = 0",
]
variables = ['A', 'K', 'C']
exo_states = ['A']
endo_states = ['K']
shocks = ['epsilon']
parameters = {
'sigma': 1.5,
'beta': 0.99,
'alpha': 0.35,
'delta': 0.025,
'rho_a': 0.9,
}
RBC = linear_dsge(
equations=equations,
variables=variables,
exo_states=exo_states,
endo_states=endo_states,
shocks=shocks,
parameters=parameters,
approximation='log_linear',
normalize={'A': 1}
)3 Once our Model RBC is defined, we can proceed with the following steps: we begin by set our initial guess for the computation of the steady-state. Since we have only 1 exo_state and using a simple guess, we can do:
RBC.set_initial_guess([1, 1])
RBC.compute_ss(method='fsolve', options={'xtol': 1e-8}) Steady-state residuals: [-1.77851329e-13 9.03721542e-14 -1.00000000e-02]
Warning: Large steady-state residuals detected.
A 1.000000
K 34.398226
C 2.589794
dtype: float644 Approximating The model around its steady state
RBC.approximate(method='analytical') (array([[-8.33790428e-03, 1.57555735e-04, 1.38972249e-01],
[ 0.00000000e+00, -1.00000000e+00, 0.00000000e+00],
[ 1.00000000e+00, 0.00000000e+00, 0.00000000e+00]]),
array([[ 0. , 0. , 0.13897225],
[-3.44974994, -1.01010101, 1. ],
[ 0.9 , 0. , 0. ]]),
array([[0.],
[0.],
[1.]]))5 Solving The model :
RBC.solve_RE_model()
print("Policy Function (f):\n", RBC.f)
print("State Transition (p):\n", RBC.p) Policy Function (f):
[[0.59492193 0.03861978]]
State Transition (p):
[[0.9 0. ]
[2.85482802 0.97148123]]6 Plotting Impulse Responses
RBC.plot_irfs()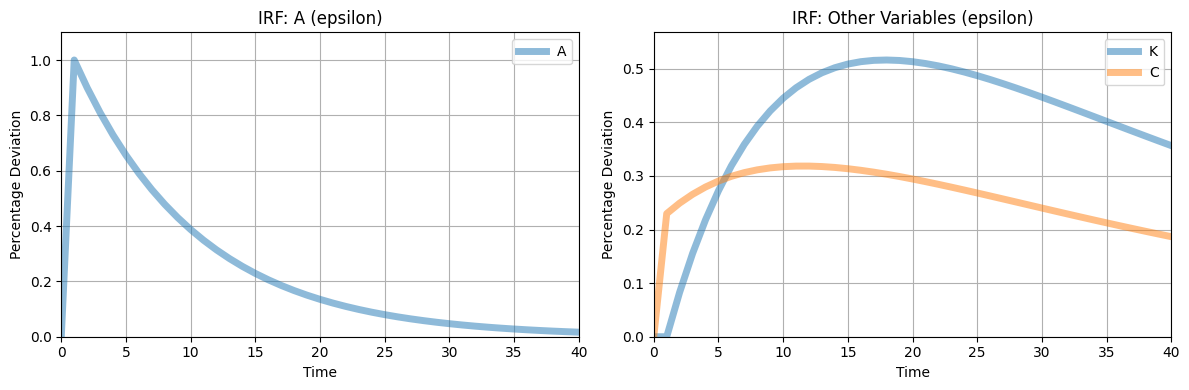
7 Simulations
RBC.simulate(T=51, drop_first=10)8 Plotting The simulations
RBC.simulations()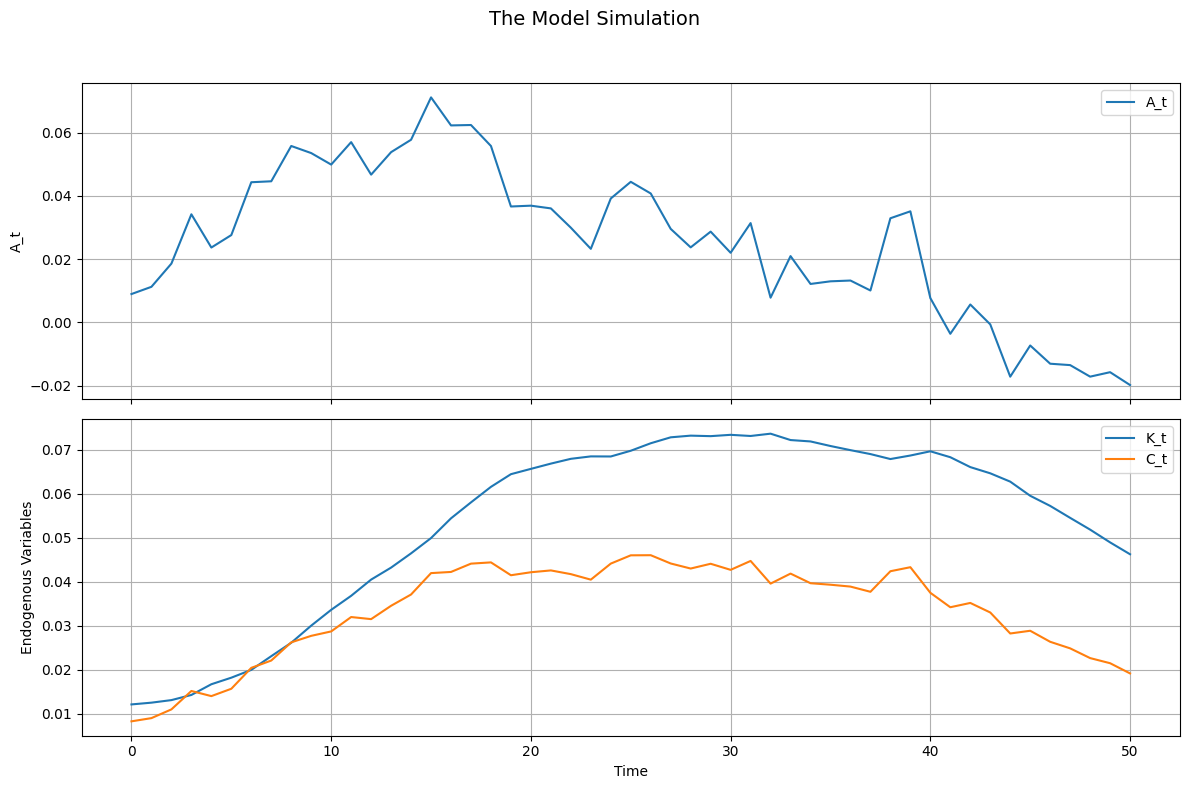
Notes
- Equation Syntax – Must use time subscripts (
_t,_tp1). - Log-Linear Approximation – Requires non-zero steady-state values.
- Shocks – If omitted, model is deterministic.
- Normalization – Overrides steady-state values for log-linear models.
Limitations
- Only
'fsolve'supported for steady states.
